 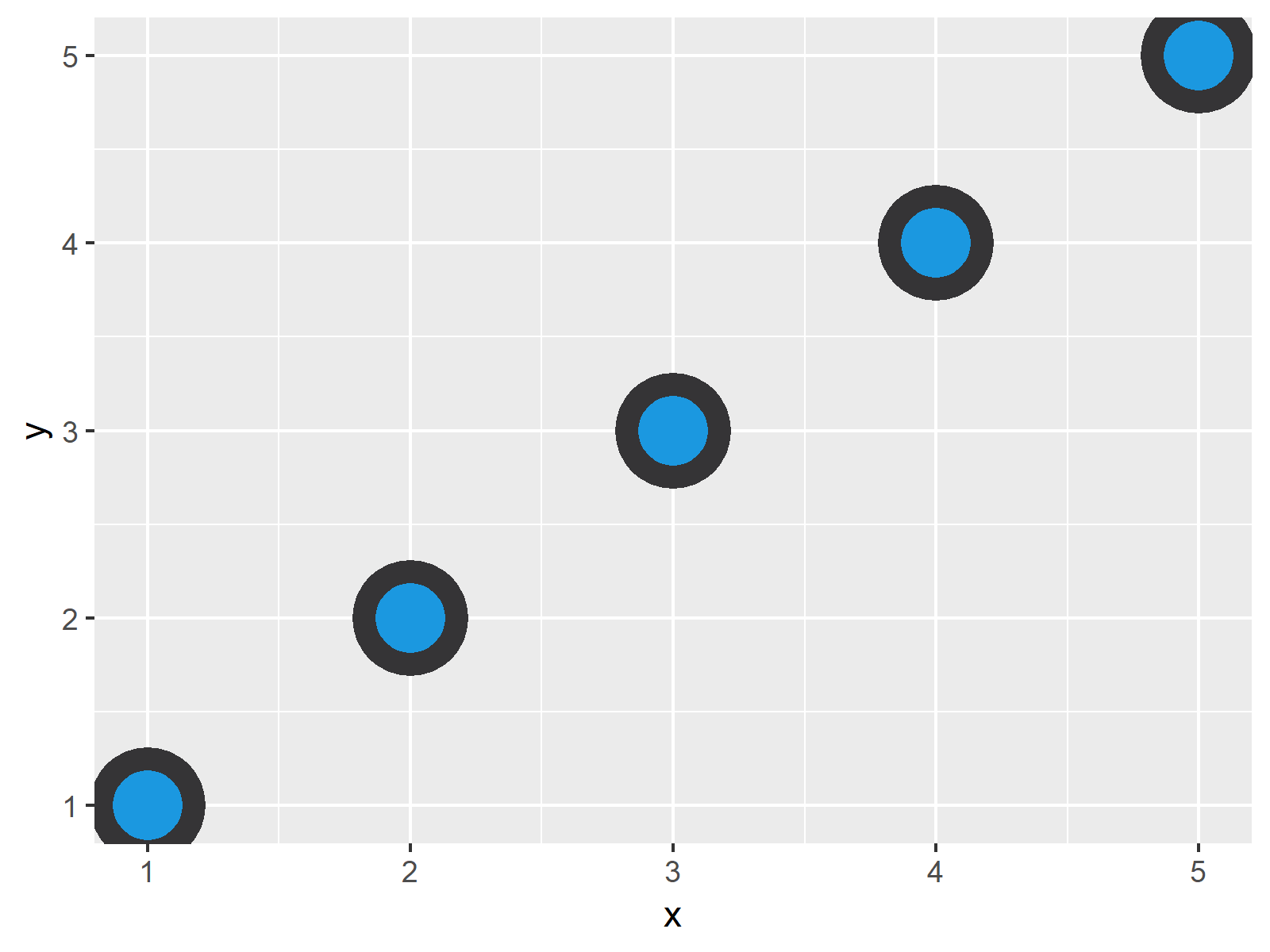
We will extend our basic template throughout this lesson as we make a variety of scatter plots and in lesson 4. Optional parameters that affect the layout and rendering, such text size, font and alignment, legend positions. the use of multiple similar subplots to look at subsets of the same data, g., linear, logarithmic, rank),Ī facet specification, i.e. One or more geometric objects that serve as the visual representations of the data, – for instance, points, lines, rectangles, contours,ĭescriptions of how the variables in the data are mapped to visual properties (aesthetics) of the geometric objects, and an associated scale (e. However, there are additional components, highlighted in bold, that can be added to our core components to enable us to generate even more diverse plot types. Second, we will use some more complicated bioinformatics data related to the RNAseq project introduced in Lesson 2.įirst, let's load our libraries using the library function, library(): See ?iris for more information about these data. These data include measurements from the petals and sepals of different Iris species including Iris setosa, versicolor, and virginica. First, we will use data available with your base R installation, the iris data set, which is stored in object iris. In this lesson we will use two different sets of data. We will do this by focusing on different types of scatter plots. The primary purpose of this lesson is to learn how to customize our ggplot2 plots. Load a different data set using new load functions Learn how to make and modify scatter plots to make fairly different overall plot representations. Learn to customize your ggplot with labels, axes, text annotations, and themes. Scatter plots and plot customization Objectives 
Practice plotting using ggplot2: Lesson 3 Practice plotting using ggplot2: Lesson 2 Lesson5: Visualizing clusters with heatmap and dendrogram Lesson 4: Stat Transformations: Bar plots, box plots, and histograms They are necessary because of the spaces in the variable name.Lesson 3: Scatter plots and ggplot2 customizationĪdd a stat to our plot with stat_ellipse().Ĭreating a publication ready volcano plot The other method, with scales, is generally a better way to do this.Īlso note the use of backticks instead of quotes. The legend title “Experimental Condtion” is long and it might look better if it were broken into two lines, but this doesn’t work very well with this method, since you would have to put a newline character in the name of the column. Ggplot ( data = pg, aes ( x = `Experimental Condition`, y = weight, fill = `Experimental Condition` )) + geom_boxplot () Manually-specified values (e.g., colors, point shapes, line types)ĭiscrete values (e.g., colors, point shapes, line types, point sizes)Ĭontinuous values (e.g., alpha, colors, point sizes)Īnother way to change the legend title and labels is to directly modify the data frame. 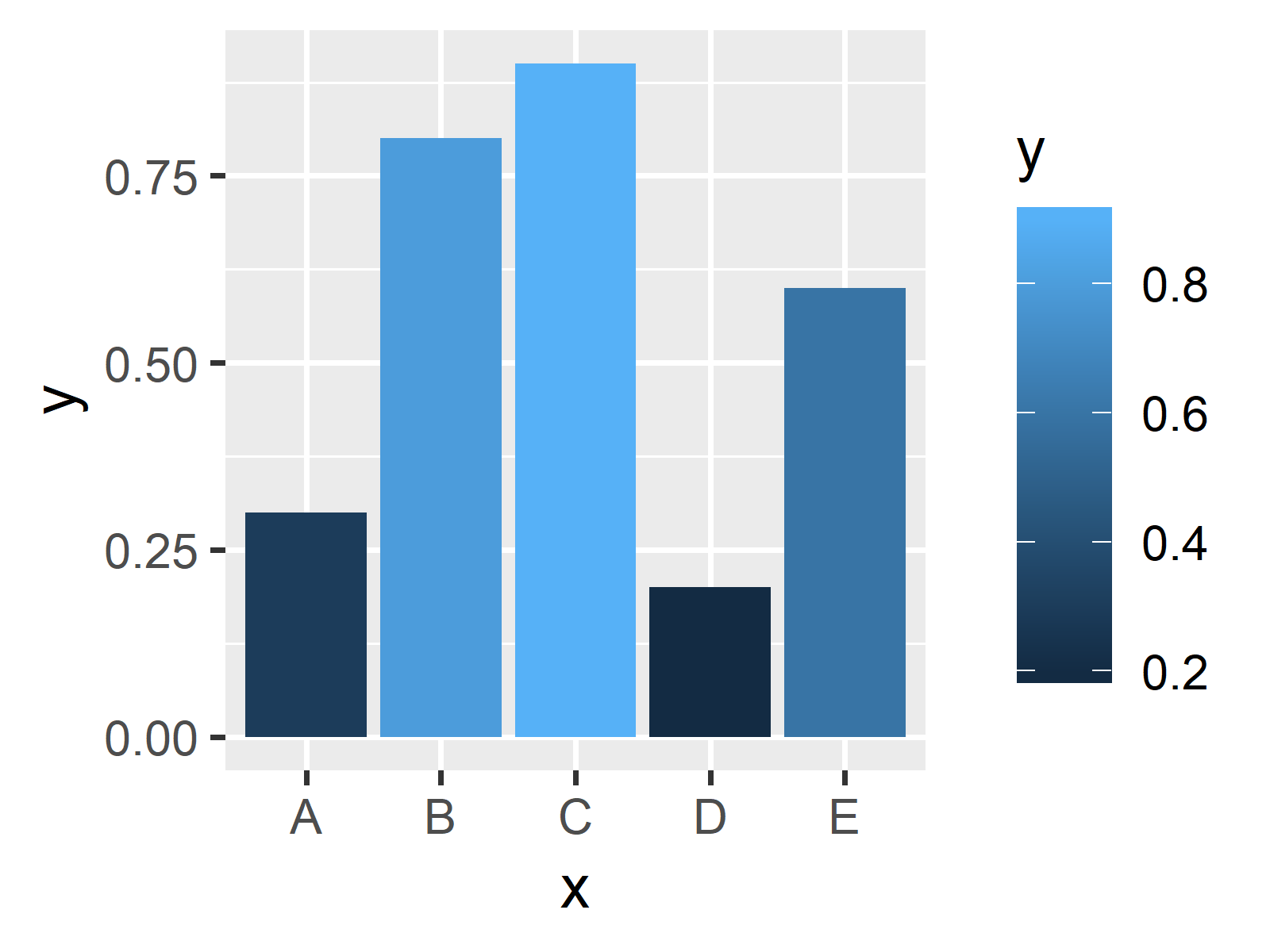
Here are some commonly-used values of xxx and yyy: xxxĮqually-spaced colors from the color wheel Lp1 + scale_colour_discrete ( name = "Payer", breaks = c ( "Female", "Male" ), labels = c ( "Woman", "Man" )) + scale_shape_discrete ( name = "Payer", breaks = c ( "Female", "Male" ), labels = c ( "Woman", "Man" )) Lp1 + scale_colour_discrete ( name = "Payer", breaks = c ( "Female", "Male" ), labels = c ( "Woman", "Man" )) # Specify both colour and shape Lp1 <- ggplot ( data = df1, aes ( x = time, y = total_bill, group = sex, shape = sex, colour = sex )) + geom_line () + geom_point () lp1 # Here's what happens if you just specify colour
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
